Rare diseases aren't rare. That's the first thing most people get wrong. Rare diseases affect over 300 million people worldwide, making them a collective public health crisis that rivals diabetes or heart disease in scale. Yet the research infrastructure supporting these patients looks nothing like it does for common conditions. Small patient populations, wildly variable symptoms, and decades of scientific neglect have left families searching for answers in a system that was never designed to find them. Disease modeling is changing that calculus entirely, giving researchers, clinicians, and biopharma partners new tools to study conditions they can't easily study any other way.
Table of Contents
- Why rare diseases require innovative modeling approaches
- Types of rare disease models: Animal, patient-derived, and in silico
- Strengths and limitations: What every stakeholder should know
- Hybrid approaches and translational impact: Why combining models matters
- The uncomfortable truth about rare disease models
- Connect with resources for rare disease research and care
- Frequently asked questions
Key Takeaways
| Point | Details |
|---|---|
| Patient scarcity solutions | Modeling overcomes limited patient numbers in rare disease research, accelerating insights and therapies. |
| Comparing modeling types | Animal, patient-derived, and in silico models each offer unique strengths for studying rare diseases. |
| Hybrid modeling advantage | Combining models improves research accuracy and speeds therapy development. |
| Validation and caution | Careful interpretation and rigorous validation are essential to avoid misleading results. |
| Accessing resources | Specialized knowledge bases help families, clinicians, and biopharma find research and treatment options. |
Why rare diseases require innovative modeling approaches
The numbers tell a stark story. There are roughly 10,000 known rare diseases, and fewer than 5% have approved treatments. Traditional drug development wasn't built for this landscape. Clinical trials need statistically significant sample sizes. Regulatory pathways assume you can recruit hundreds of patients. Neither assumption holds when a disease affects 50 families worldwide.
"Rare diseases collectively affect over 300 million people, but small patient populations, limited data, and heterogeneous phenotypes hinder traditional research and drug development." — Nature Digital Medicine
This is exactly why rare disease research needs modeling approaches that don't depend on large patient cohorts. Models let scientists recreate disease biology in a controlled environment, observe how a disease progresses, and test candidate treatments without waiting years for natural history data to accumulate. They're not a workaround. They're the only viable path forward for most ultra-rare conditions.
The core challenges that modeling addresses include:
- Patient scarcity: A disease affecting 200 people globally cannot support a traditional randomized controlled trial
- Phenotypic diversity: Two patients with the same genetic mutation can present completely differently, making population-level data misleading
- Disease mechanism gaps: For many rare diseases, the underlying biology is simply unknown, making rational drug design impossible without a model to study
- Ethical constraints: You can't withhold treatment from dying patients to run a placebo arm when no treatment exists
- Time pressure: Families can't wait 15 years for a drug to clear traditional development pipelines
Models solve each of these problems in different ways, depending on what type of model you use.
Types of rare disease models: Animal, patient-derived, and in silico
Not all models are created equal. Each major approach has a different purpose, a different level of biological fidelity, and a different role in the research pipeline.
| Model type | Biological fidelity | Scalability | Cost | Best use case |
|---|---|---|---|---|
| Animal models | Moderate to high | Moderate | High | Mechanism discovery, in vivo testing |
| Patient-derived (iPSC/organoids) | Very high | Limited | Very high | Drug screening, genotype-phenotype correlation |
| In silico | Variable | Very high | Low to moderate | Virtual trials, dosing prediction, data synthesis |
Animal models have anchored rare disease research for decades. Genetically engineered mice, rats, and zebrafish are crucial for understanding disease pathogenesis, testing therapies at a whole-organism level, and bridging the gap when human studies aren't ethically feasible. Zebrafish in particular have become valuable because their embryos are transparent, allowing researchers to watch organ development in real time. Mouse models expressing specific gene mutations have generated foundational insights into conditions like Duchenne muscular dystrophy, cystic fibrosis, and dozens of lysosomal storage disorders.

Patient-derived models represent a genuine leap forward. When a lab reprograms a patient's own skin or blood cells into induced pluripotent stem cells (iPSCs) and then differentiates those cells into neurons, cardiomyocytes, or hepatocytes, it creates a living replica of that patient's disease biology. iPSC models and organoids recapitulate human-specific genotype-phenotype correlations, disease progression, and enable drug screening at a level no animal model can match. A mouse with a KCNQ2 mutation doesn't have a human brain. A dish of neurons derived from a child with that mutation does.
In silico models take a different approach entirely. Instead of growing cells or breeding animals, they use computational algorithms, mathematical equations, and machine learning to simulate how a disease behaves and how a drug might interact with it. In silico models simulate mechanisms and drug responses, address data scarcity, and support precision dosing in ways that wet-lab experiments simply can't achieve at scale. For an ultra-rare disease with 30 known patients, an in silico model can generate thousands of virtual patient profiles based on known biology, giving researchers a synthetic dataset to work with.
Key advantages of patient-derived models that make them especially valuable in rare disease contexts:
- They carry the exact genetic variant responsible for the patient's disease
- They can model cell types directly affected by the condition, such as neurons for neurological diseases or cardiomyocytes for cardiac channelopathies
- They allow drug screens across hundreds of compounds simultaneously
- They can be used to test personalized antisense oligonucleotides (ASOs) designed for a specific patient's mutation
Pro Tip: If you're a clinician evaluating modeling options for a patient with an ultra-rare disease, prioritize iPSC modeling when the disease primarily affects a specific cell type that can be reliably differentiated. In silico tools work best when you already have some mechanistic understanding of the disease pathway to build from.
Strengths and limitations: What every stakeholder should know
Understanding what each model does well is only half the picture. Knowing where each model breaks down protects against over-interpreting results and wasting precious time chasing leads that won't translate to patients.
| Approach | Key strengths | Core limitations |
|---|---|---|
| Animal models | In vivo pharmacokinetics, whole-organism toxicity, mechanism validation | Species-specific biology often doesn't translate; models underestimate prevalence without diverse genetic data |
| Patient-derived | High human relevance, personalized mutation modeling | Limited scalability, high cost, technically demanding, 2D cultures miss organ-level complexity |
| In silico | Scalable, fast, integrates heterogeneous datasets | Data quality and validation gaps, interpretability challenges, requires experimental confirmation |
The animal model problem deserves more attention than it typically gets. Mouse models have driven enormous scientific progress, but there's a growing recognition that they regularly fail to capture human pathophysiology. Inflammatory pathways differ. Drug metabolism genes differ. Brain structure and connectivity differ in ways that matter enormously when you're studying a rare neurological condition. Animal models often fail to fully capture human disease biology, and genetic models specifically tend to underestimate how prevalent a condition is when they're built from non-diverse genetic data.
For patient-derived models, the scalability wall is real. Creating an iPSC line from a single patient takes months of specialized work and significant cost. Differentiating those cells into a specific cell type adds more time. Running a drug screen on top of that requires infrastructure most academic labs don't have. This is not a workflow you can run for 500 compounds without serious resources.
Here's a clear framework for navigating model selection:
- Start with in silico to generate hypotheses and prioritize candidate compounds when you have limited resources and some existing biological data
- Move to patient-derived models to validate promising candidates in human-relevant biology and identify patient-specific response patterns
- Use animal models to confirm efficacy in a whole-organism context before clinical translation, particularly for safety and pharmacokinetics
- Validate across all three before making major resource commitments or clinical decisions
Pro Tip: When reviewing modeling data in a disease treatment search, always ask how the model was validated and whether results were replicated in at least one other system. A single model showing a positive result is a hypothesis, not a finding.
In silico models carry their own risks. They are only as good as the data they're trained on, and for rare diseases, that data is thin. A model trained on a handful of published case reports can produce confident-looking predictions that are badly wrong because the training data wasn't representative. Interpretability is another issue: black-box machine learning models may identify patterns that researchers can't mechanistically explain, making regulatory conversations difficult.
Organoids, which are three-dimensional tissue structures grown from iPSCs, address some limitations of flat cell cultures but introduce new ones. They're expensive to grow, difficult to standardize, and hard to scale. Vascularization remains an unsolved problem, meaning most organoids lack the blood supply that real organs depend on, which limits how well they represent drug distribution in a living system.
Hybrid approaches and translational impact: Why combining models matters
The most sophisticated teams working on rare diseases today don't pick one model type and commit to it. They combine all three, using each where it's strongest and checking results against the others.
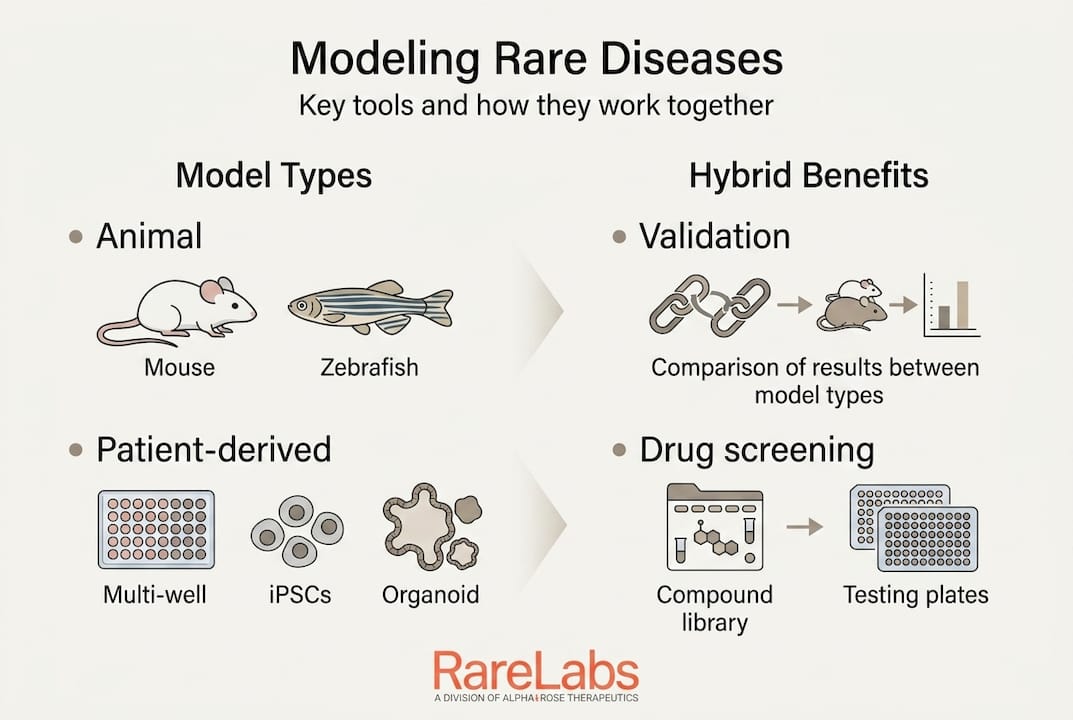
Hybrid approaches combining in silico predictions with patient-derived organoids and animal validation offer the most promise for rare disease research, addressing individual model limitations while amplifying their collective strengths for translation. This isn't just an academic preference. It's increasingly what regulatory bodies expect to see in submissions for rare disease treatments.
The practical benefits of a hybrid modeling strategy include:
- Higher predictive accuracy: Results that hold across three different model types are far more likely to translate to clinical benefit
- Faster screening: In silico tools can shortlist 10,000 compounds to 50 candidates that then get tested in patient-derived cells, rather than running every compound in expensive biological assays
- De-risked development: Animal validation before clinical trials catches safety signals early, reducing the probability of costly late-stage failures
- Better regulatory packages: The FDA's rare disease evidence principles explicitly recognize that non-traditional evidence, including modeling data, can support approval decisions when conventional trials aren't feasible
For biopharma partners, models de-risk drug development and enable drug repurposing, preclinical evaluation, and evidence generation for ultra-rare diseases via the FDA's Rare Disease Evidence Principles framework. This matters enormously for commercial strategy. A compound that failed a Phase 2 trial in a common indication might be exactly the right fit for a rare disease population, and a patient-derived model can identify that match in months rather than years.
Key insight: For every 10 compounds that enter preclinical testing, only 1 typically reaches patients. Hybrid modeling approaches are steadily improving that ratio for rare disease treatment strategies, by catching the wrong candidates earlier and surfacing the right ones faster.
The translational payoff is real. Programs that integrate patient-derived validation into their screening pipeline consistently produce stronger clinical candidates because they're testing in biology that actually reflects the patients they're trying to treat. That's not a philosophical preference. It's a practical efficiency gain.
The uncomfortable truth about rare disease models
Here's what most overviews get wrong: they present modeling as a solved problem, a clean toolkit you can pick from and trust. The reality is more complicated, and patients and families deserve to hear it plainly.
Models are powerful. They're also imperfect in ways that can mislead researchers who don't interrogate their assumptions. The field still lacks standardized validation frameworks for patient-derived models. Two labs running drug screens on iPSC-derived neurons from the same genetic background can produce meaningfully different results depending on their differentiation protocols, passage numbers, and quality controls. Until those standards are established and enforced, results need to be interpreted with caution.
Models must be cautiously interpreted due to species differences, data biases, and validation gaps. Regulatory alignment and open-source data sharing are what will actually move the needle on clinical impact. This is where the rare disease field has real room to grow. Proprietary data silos slow progress dramatically. A family fighting a condition affecting 40 people worldwide doesn't have time for institutions to protect competitive advantage. Open data sharing, preregistered modeling protocols, and early regulatory engagement aren't idealistic goals. They're practical necessities.
Patients and families should ask hard questions when they hear that a modeling study showed a promising result. Ask whether the model was validated against real patient biology. Ask whether the result has been replicated. Ask whether the lab used a patient-derived model or an animal model or a cell line, and understand what that means for how much confidence to place in the finding. Our rare disease expertise is built around rigorous, transparent modeling precisely because these questions matter so much when families are making decisions under pressure.
The path forward requires patients, clinicians, and biopharma partners working together with shared data and honest communication about uncertainty. Models are the best tools we have. They're not oracles, and treating them as such is the fastest way to stall progress.
Connect with resources for rare disease research and care
Navigating rare disease research can feel like searching for a light switch in a dark room. Families deserve better than a literature search and a prayer.
RareLabs builds patient-specific disease models using iPSCs, CRISPR gene editing, and parallel drug screening to accelerate the search for viable treatments. Whether you're a patient, a clinician, or a biopharma partner, our rare disease knowledge base offers research insights, process explanations, and scientific context to help you understand what's possible. If you're actively looking for treatment options for a patient with an undiagnosed or ultra-rare condition, our treatment search for rare diseases is built to move fast, stay rigorous, and put the patient's own biology at the center of every decision.
Frequently asked questions
How do animal models help study rare diseases?
Animal models like mice and zebrafish allow researchers to uncover disease mechanisms and test therapies in a whole-organism context when human trials aren't feasible, and they remain essential for safety and pharmacokinetic data before any clinical translation. Animal models are crucial for elucidating pathogenesis and bridging gaps where human studies are limited.
What are in silico models in rare disease research?
In silico models use computational simulations and machine learning to predict disease progression and drug responses, which is especially valuable for rare diseases where patient data is scarce. In silico models simulate disease mechanisms and predict drug responses, enabling virtual trials that would be impossible with a small patient population.
What are the main limitations in current rare disease models?
Every major modeling approach faces constraints including species-specific biology gaps in animal models, scalability and standardization issues in patient-derived models, and data quality and validation gaps in in silico tools. In silico limitations include data quality issues, lack of validation standards, interpretability problems, and the need for hybrid integration to produce reliable, clinically relevant results.
Why combine different modeling approaches?
No single model type captures the full picture of a rare disease, so combining animal, patient-derived, and in silico methods produces more accurate, reproducible results and reduces the risk of chasing false leads into expensive clinical programs. Hybrid approaches combining in silico predictions with patient-derived organoids and animal validation offer the strongest translational outcomes for rare disease drug development.
